In a study published in Cell Reports, BI Changhao's lab and ZHANG Xueli's lab at Tianjin Institute of Industrial Biotechnology of the Chinese Academy of Sciences constructed enhanced GBE variants for wider application.
The optimized GBEs were obtained via integrating transactivation modules, SunTag-system and Cas9 variant.
Firstly, the transactivation modules were selected to fused to GBE, and the higher efficiency was obtained at all testing loci in HEK293T cells. In addition, the SunTag system was utilized to optimize the performance of GBE. The higher efficiency, higher purity and extended editing window were acquired. Finally, the SpRY-Cas9 was fused to GBE to construct the SpRY-GBE variant, which could recognize the non-NGG PAM and provide the wider targeting scope for GBE.
The GBE variants with superior performance were acquired, which provides wider application of GBE for targeting more genomic sites with higher efficiency and specificity, thus improving the potential for the genetic therapy of more mutational related diseases
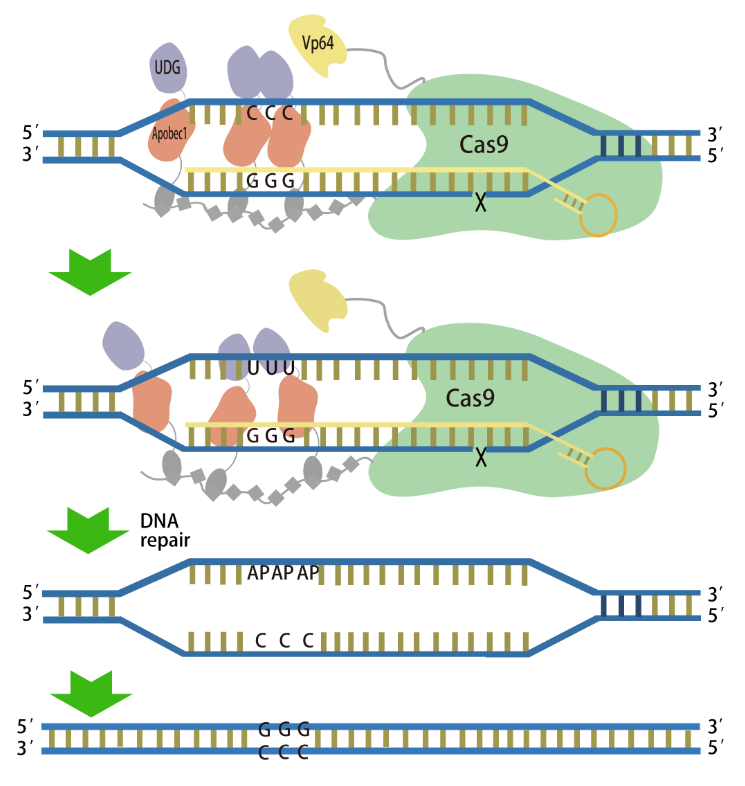
Working diagram of the SunTag-GBEs ( Image by TIBCAS)
Contact:
Prof. BI Changhao
Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences.
Phone: +8618302294563
Email: bi_ch@tib.cas.cn