Methanol is an energy-rich and readily available feedstock for biomanufacturing. Microbial metabolism of methanol is expected to provide more reducing power than sugars and consequently lead to higher product yield. Recently, massive efforts have been devoted to construction of synthetic methylotrophs, which represent engineered platform microorganisms with the capability to incorporate methanol for building up cellular constituents and producing chemicals. However, the existing synthetic methylotrophs still suffer from low methanol bioconversion efficiency, which hinders the development of methanol-based biomanufacturing.
Recently, researchers from Laboratory of Systems & Synthetic Biotechnology led by Prof. ZHENG Ping and Systems Biology Center led by Prof. SUN Jibin at Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences reviewed the ongoing efforts to construct and optimize synthetic methylotrophs for methanol-based biomanufacturing. Enzyme and pathway engineering strategies such as discovering and engineering key enzymes, enhancing supply of formaldehyde acceptors, and coupling methanol metabolism and cell growth are now commonly used to improve synthetic methylotrophy. The low tolerance to methanol was a previously ignored issue for engineering synthetic methylotrophs. The same group further applied adaptive laboratory evolution and tolerance engineering strategies to improve the methanol tolerance and conversion in synthetic methylotrophic Corynebacterium glutamicum. The evolved mutants can tolerate higher concentrations of methanol and show improved growth and methanol utilization. Methanol and co-substrate xylose were utilized with a molar ratio over 7:1, suggesting methanol was the major carbon source of the synthetic methylotroph. Genome and transcriptome analyses and reverse metabolic engineering were then conducted to study the mechanism of improved methanol tolerance and bioconversion. It was found that increased methanol concentrations rebalanced methylotrophic metabolism by down-regulating glycolysis and upregulating amino acid biosynthesis, oxidative phosphorylation, ribosome biosynthesis, and parts of TCA cycle (as shown in the following figure). A mutation in the O-acetyl-L-homoserine sulfhydrylase Cgl0653 was proven crucial for methanol tolerance, which decreased the formation of L-methionine analog in the presence of methanol. Another mutation in the methanol-induced membrane-bound transporter Cgl0833 also contributed largely to methanol tolerance, whereas the mechanism was still unclear. This study underlined the importance of tolerance engineering in developing superior synthetic methylotrophs and laid foundation for future optimizing synthetic methylotrophy.
These studies were supported by National Key R&D Program of China (2018YFA0901500), National Natural Science Foundation of China (31700044), etc. The review entitled “Synthetic methylotrophy: a practical solution for methanol-based biomanufacturing” has been published in Trends in Biotechnology. The research paper entitled “Adaptive laboratory evolution enhances methanol tolerance and conversion in engineeredCorynebacterium glutamicum” has been published in Communications Biology. Associate professor Dr. WANG Yu is the first author of these two papers. Prof. ZHENG Ping and Prof. SUN Jibin are the corresponding authors.
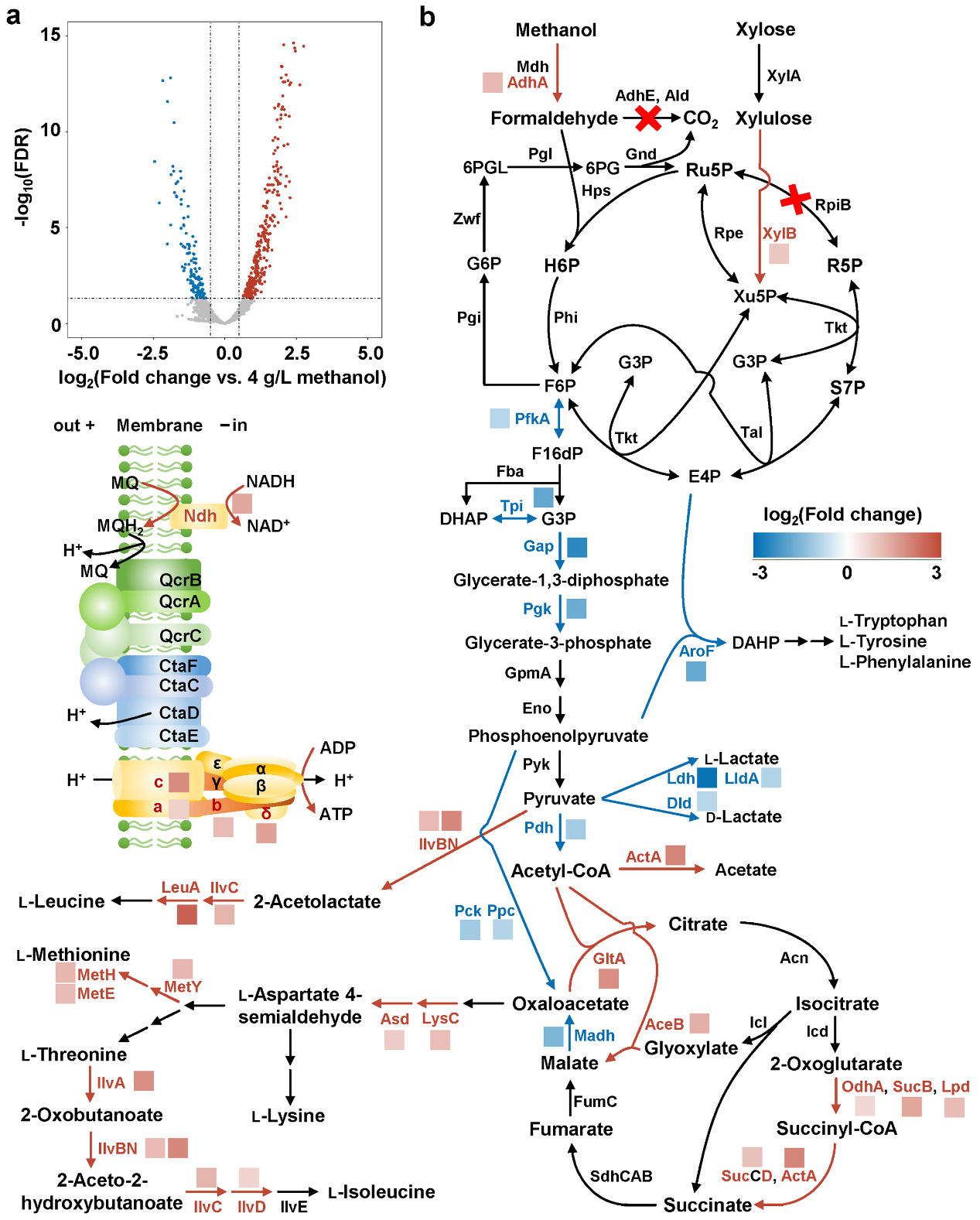
Transcriptome analysis of a methylotrophic C. glutamicum strain cultivated with different concentrations of methanol. a Differential transcript levels as determined by RNA sequencing (15 g/L vs. 4 g/L methanol). b Gene transcript level changes related to central metabolism pathways and oxidative phosphorylation (15 g/L vs. 4 g/L methanol) (Image by Prof. ZHENG Ping’s group)
Contact:Prof. ZHENG Ping
Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences
Phone: 0086-22-84861943
Email: zheng_p@tib.cas.cn