Metabolomics has been a potential tool for strain improvement through analyzing metabolite changes in the context of different conditions. However, the availability of a universal metabolite profiling analysis is still a big challenge.
Recently, the research group led by Prof. Zheng Ping and Prof. Sun Jibin presented an optimized liquid chromatography-tandem mass spectrometry-based metabolomics methodology for Corynebacterium glutamicum, an important industrial workhorse. It was found that quenching the cellular metabolism with 5-fold volume of -20°C 40% methanol was highly recommended due to its lower cell damage rate and higher intracellular metabolites recovery rate. For extracting intracellular metabolites, ethanol/water (3:1, v/v) at 100°C combined with acidic acetonitrile/water (1:1, v/v, with 0.1% formic acid) at -20°C achieved the unbiased metabolite profiling of C. glutamicum. The established methodology was then applied to investigate the intracellular metabolite differences between C. glutamicum ATCC 13032 and an mscCG-deleted mutant under biotin limitation condition. The optimized metabolomics methodology holds promise for promoting studies on metabolic mechanism of C. glutamicum.
The work entitled “Comprehensive optimization of the metabolomic methodology for metabolite profiling ofCorynebacterium glutamicum” has been published in Applied Microbiology and Biotechnology . M.S. Zhang Qiongqiong and Dr. Zheng Xiaomei of TIB are co-first authors of this paper. This study was supported by the National Natural Sciences Foundation of China, Chinese Academy of Sciences Key Project.
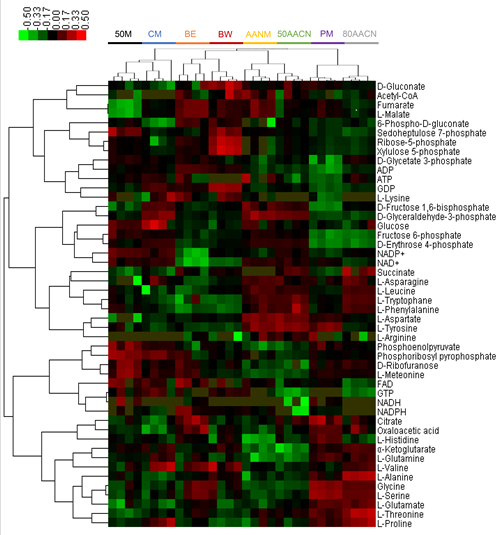
Fig. 1 Hierarchical clustering analysis of intracellular metabolites from C. glutamicum ATCC 13032 extracted by eight extraction solutions.
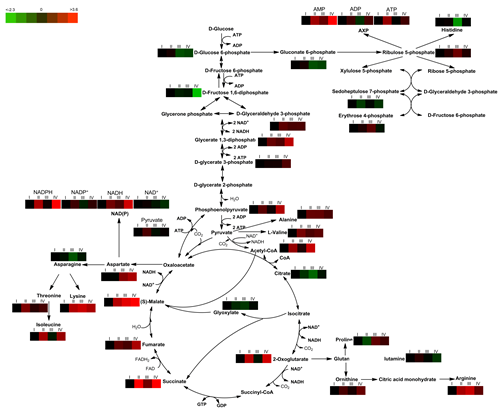
Fig. 2 Intracellular metabolite profiling of intermediates involved in central metabolic pathway in C. glutamicum ATCC 13032 and ATCC 13032ΔmscCG under biotin-limited condition.
Contact:Prof. ZHENG Ping
Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences.
Email: zheng_p@tib.cas.cn